Artifact Removal & Biomarker Segmentation
Overview
EECE571T - Advanced Machine Learning Tools, is a graduate level machine learning course I took at UBC. A large part of this course was the final project for which I choose to do artifact removal and biomarker segmentation of FOXP3+ biomarkers for follicular lymphoma TMA cores in conjunction with the British Columbia Cancer Agency.
The purpose of the project was to introduce a quantitative method of evaluating FOXP3+ biomarker counts in TMA cores, and improve upon industry standard - usually estimated by eye by a Pathologist or by the software Aperio.
One major obstacle was the frequent presence of artifacts in the cores which would completely overpower the actual positive biomarkers themselves. These had to be ignored by Pathologists, and removed by hand in Aperio.
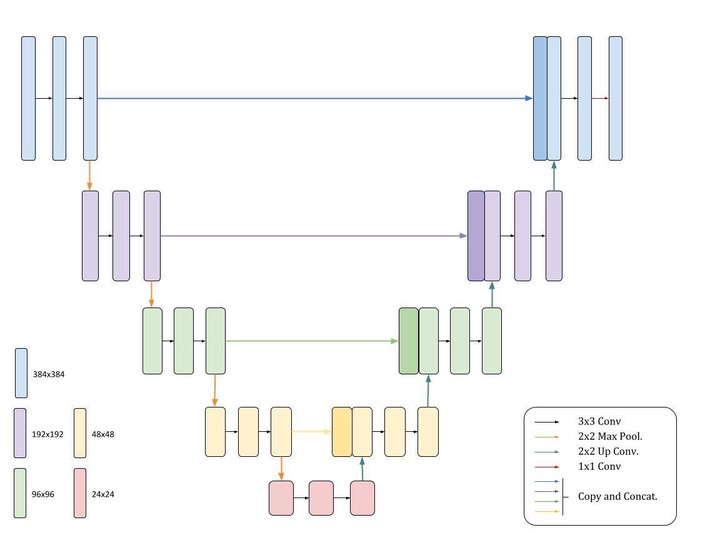
As such, the proposed framework is broken into artifact segmentation, to segment and remove artifacts, and marker segmentation to identify the biomarkers. In both cases, the input images were very large ($2886\times 2886$), so to preserve global and local structure, patches were made and fed into UNets to produce binary masks for both artifacts and markers. These methods and results are discussed in detail in the final report paper.